Survival analysis is a statistical technique used to analyze time-to-event data, such as the time until death or the time until the failure of a machine. R is a popular programming language used by statisticians and data analysts for data analysis, visualization, and modeling.
In R, survival analysis can be performed using the survival package. This package provides functions for fitting different types of survival models and for conducting various types of survival analyses, such as Kaplan-Meier curves, Cox proportional hazards regression, and parametric survival models.

Download:
To begin, you will need to load the survival package into R by typing:
library(survival)
The first step in survival analysis is to create a survival object. A survival object is a data structure that contains information about the time-to-event data, including the time-to-event (often called “survival time”), the event status (often called “censoring status”), and any covariates that may affect the survival time.
To create a survival object, you can use the Surv() function. For example, suppose you have a dataset called mydata that contains information on the survival time and censoring status of patients in a clinical trial. You can create a survival object as follows:
my.survival <- Surv(time = mydata$time, event = mydata$status)
In this example, time is a vector of survival times, and status is a vector of censoring statuses (0 if the event was censored, 1 if the event occurred). The Surv() function combines these vectors into a single survival object.
Once you have created a survival object, you can use it to fit survival models. The most commonly used survival model is the Cox proportional hazards regression model, which allows you to estimate the effect of covariates on the hazard rate (i.e., the instantaneous risk of experiencing the event at any given time). To fit a Cox proportional hazards model in R, you can use the coxph() function. For example:
my.coxph <- coxph(formula = Surv(time, status) ~ covariate1 + covariate2, data = mydata)
In this example, formula is a formula that specifies the survival object and the covariates to be included in the model, and data is the name of the dataset containing the variables. The output of the coxph() function is an object of class “coxph”, which can be used to obtain estimates of the hazard ratio (i.e., the relative hazard of experiencing the event associated with a one-unit increase in a covariate) and other model parameters.
In addition to Cox proportional hazards regression, there are many other types of survival models that can be fitted using the survival package, such as parametric survival models, accelerated failure time models, and frailty models. The package also provides functions for conducting various types of survival analyses, such as Kaplan-Meier curves and log-rank tests.
Overall, survival analysis is a powerful method for analyzing time-to-event data in R, and the survival package provides a wide range of functions and tools for conducting different types of survival analyses.
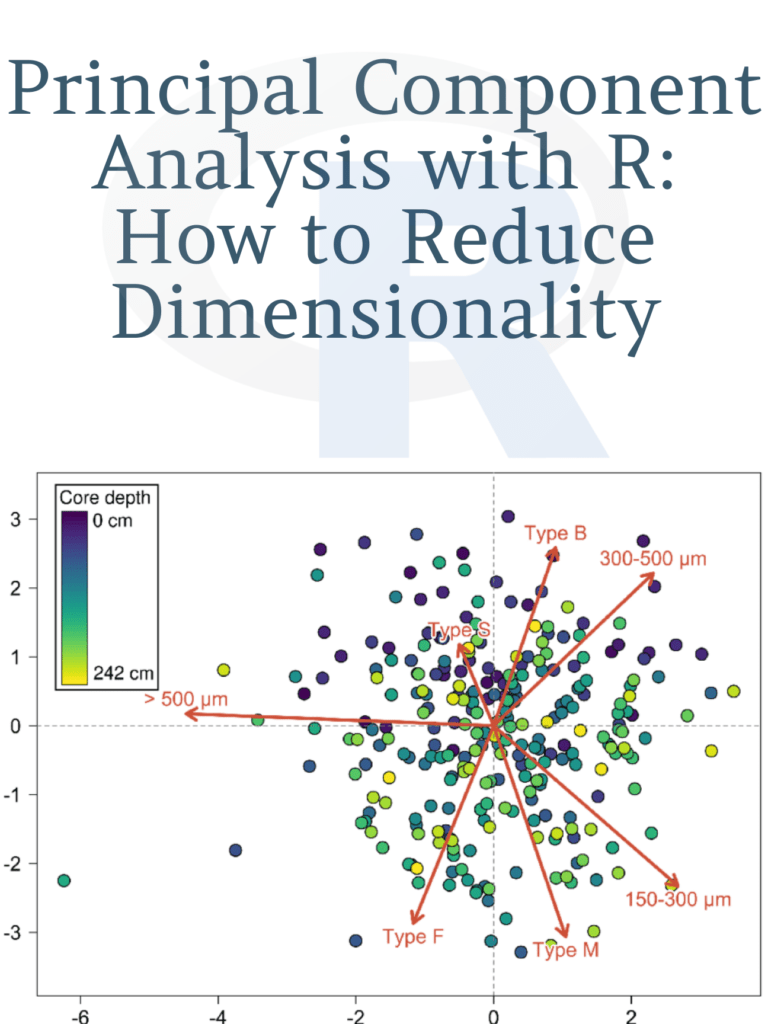
Learn: Principal Component Analysis with R: How to Reduce Dimensionality