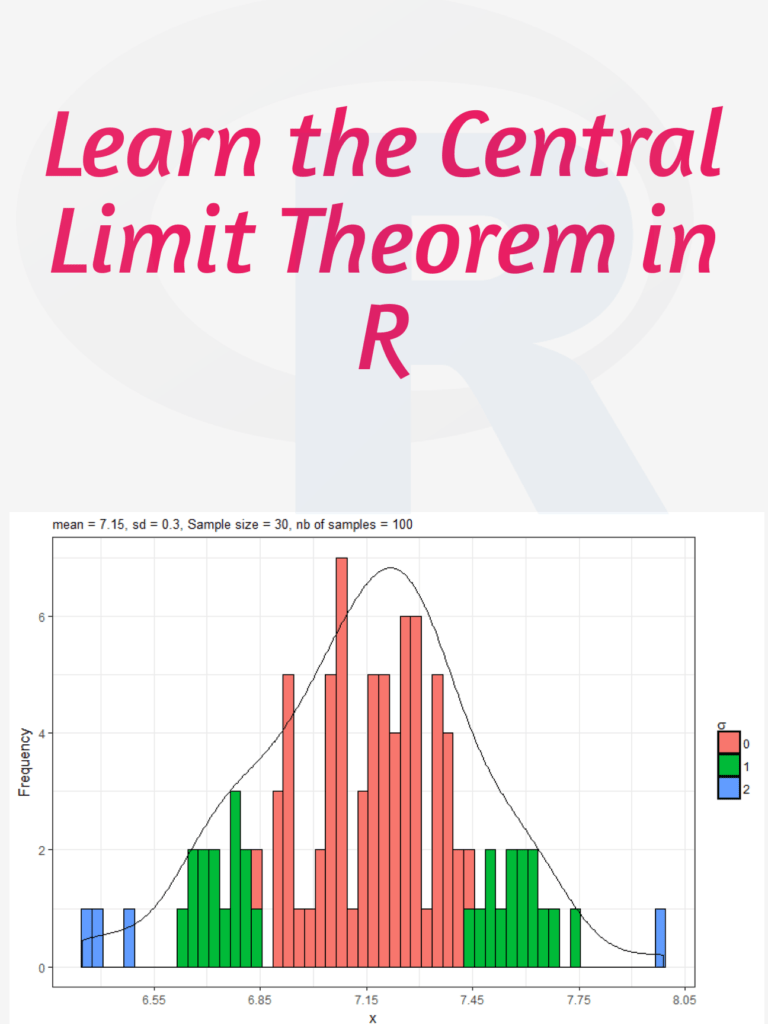
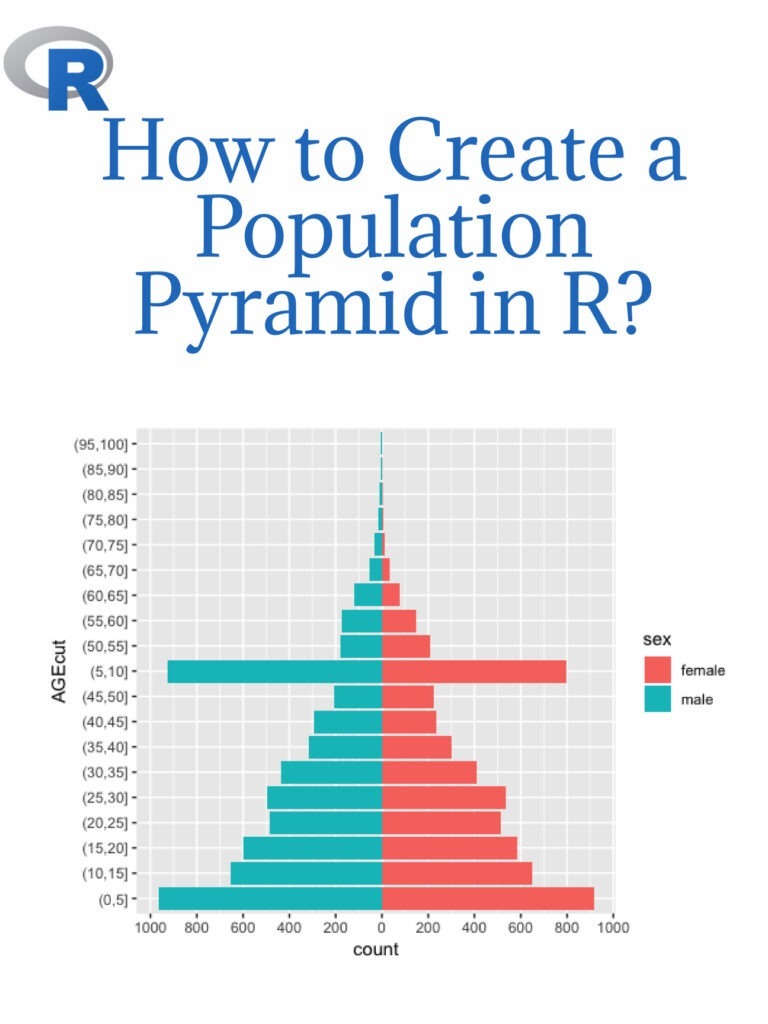
Creating a normal distribution plot using ggplot2 in R: The normal distribution is a probability distribution that is often used to model real-world phenomena, such as the distribution of test scores or the heights of a population. It is a bell-shaped curve that is symmetric around its mean value, and its standard deviation determines its spread. In this article, we will walk through the steps of creating a normal distribution plot using the ggplot2 package in R.

Download:
Step 1: Generate a dataset
To create a normal distribution plot, we first need to generate a dataset that follows a normal distribution. We can use the rnorm function in R to generate a random sample of numbers that follow a normal distribution with a specified mean and standard deviation. For example, let’s generate a sample of 1000 numbers with a mean of 50 and a standard deviation of 10:
set.seed(123) # for reproducibility
data <- data.frame(x = rnorm(1000, mean = 50, sd = 10))
This will create a data frame with one column, “x”, that contains our randomly generated numbers.
Step 2: Create a histogram
Next, we can create a histogram of our data using the ggplot2 package. A histogram is a graphical representation of the distribution of a dataset, and it can help us visualize the shape of our normal distribution.
library(ggplot2)
ggplot(data, aes(x = x)) +
geom_histogram(binwidth = 1, color = "black", fill = "white") +
labs(x = "Values", y = "Frequency", title = "Histogram of Normal Distribution")
This code will create a histogram with a binwidth of 1, a black border, and white fill. The x-axis will be labeled “Values”, the y-axis will be labeled “Frequency”, and the title of the plot will be “Histogram of Normal Distribution”.
Step 3: Add a density curve
To make our plot more informative, we can add a density curve to show the shape of the normal distribution. A density curve is a smoothed version of the histogram that shows the distribution of our data more clearly.
ggplot(data, aes(x = x)) +
geom_histogram(binwidth = 1, color = "black", fill = "white") +
geom_density(color = "blue", size = 1) +
labs(x = "Values", y = "Density", title = "Histogram and Density Curve of Normal Distribution")
This code will add a blue density curve to our histogram with a size of 1. The x-axis will be labeled “Values”, the y-axis will be labeled “Density”, and the title of the plot will be “Histogram and Density Curve of Normal Distribution”.
Step 4: Customize the plot
Finally, we can customize our plot by adding axis labels, changing the colors and fonts, and adjusting the layout.
ggplot(data, aes(x = x)) +
geom_histogram(binwidth = 1, color = "black", fill = "#69b3a2") +
geom_density(color = "#e9c46a", size = 1) +
labs(x = "Values", y = "Density", title = "Normal Distribution Plot") +
theme_minimal() +
theme(plot.title = element_text(size = 18, face = "bold"),
axis.title = element_text(size = 14, face = "bold"),
axis.text = element_text(size = 12),
legend.position = "none")
This code will change the fill color of the histogram to “#69b3a2” and the color of the density curve to “#e9c46